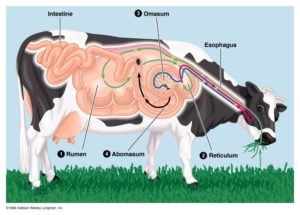
Grazing animals like cattle, buffalo, sheep, goat have four-chambered stomach called as rumen. It contains a complex bionetwork, where all types of microbes including bacteria, fungi, bacteriophages, archaea and protozoa exist in close proximity. Some of these microbes interact with each other in a synergistic relationship to extract energy by breaking lignocellulosic bond with help of lignocellulolytic enzymes like cellulose to support digestion of the host. This yields volatile fatty acids (VFAs, acetate, butyrate, propionate), formic acid, H2CO3 and CH4. With increase in global warming animals also contribute to increase in environmental temperature by releasing methane into the environment. Methane has 20 times more global warming potential than CO2, therefore, without rumen microorganisms, or when the rumen ecosystem is disturbed, there can be adverse effects on animal health and productivity and on environment as well. Now a days to reduce methane emissions by altering the rumen ecosystem is scientific challenge. Understanding of complex microbial communities’ function and microbial interaction within their niches represents a major challenge for rumen microbiologists even today. Advancing molecular biology and genomics offer new opportunities to conduct a holistic examination of the structure and function of rumen microbial communities. Analyzing the role of individual group of microbes, microbial structure, composition, and interaction may provide functional dimensions of the rumen network that will help to achieve a major goal of rumen microbiology.
Rumen Microbial Ecosystem
Rumen consist of anaerobic environment with microbes that helps in digestion of fibrous feed. Rumen microbes are divided into different groups. Bacteria (bacteria), Archaea (methanogens), and Eukarya (protozoa and fungi). Rumen microbes produces enzyme cellulose that helps in digestion of plant structural carbohydrates such as lignin, cellulose, and hemicellulose,which other animal can’t. Since strict anaerobic conditions (−150 to −350 mV) prevail in the rumen, the natural rumen microflora consists of strict anaerobes. The rumen is open to the external environment, and there is a continuous flow of material into and out of the rumen. It has a dry matter content of 10–18%, and temperature is typically regulated between 38 and 41°C. Rumen has motility 3/min. The physiological pH range is between 5.5 and 6.9, and it is one of the most variable factors in the rumen environment. Cattle has 58 liter rumen fluid volume (15-21% of body wt). It varies with individual animal consuming more feed have more rumen fluid volume. Rumen volume is greater with roughages. However, the buffering capacity of the rumen content is quite high.
| Physical, chemical, and microbiological characteristics of rumen ecosystem | ||
| Physical properties | Dry matter (%) | 110–18 |
| Osmolality | 250–350 mOsmol/Kg | |
| pH | 5.5–7.2 (Mean 6.4) | |
| Redox potential | −350 to −400 mV | |
| Temperature | 38–41 °C | |
| Chemical properties
|
Amino acids and oligopeptides | <1 mmol/L present 2–3 h post feeding |
| Ammonia | 2–12 mmol/L | |
| Dietary (cellulose, hemicelluloses, pectin) component | Always present
|
|
| Endogenous (mucopolysaccharides) | Always present | |
| Gas phase (%) | CO2 65%; CH4 27%, N2 7%; O2 0.6%, H2 0.2% | |
| Microbiological properties | Bacteria | 1010–11 per g (>200 species) |
| Ciliate protozoa | 10 4–6 per g (25 genera) | |
| Anaerobic fungi | 10 3–5 per g (6 genera) | |
| Bacteriophage 1 | 10 7–9 per g particles per ml | |
Source: Adapted from Mackie et al. (1999), Castro-Montoya et al. (2011)
Bacteria
Rumen contains different types of bacteria, which are most actively involved in the plant fiber degradation, as revealed by the fact that bacteria associated with feed particles account for nearly 50–75 % of the total microbial population. Majority of the rumen bacteria are obligate anaerobes. The adhesion of microbes solid substances is an important factor in successful competition and survival in the rumen, as well as in the digestion of solid feed. In addition, most of the rumen endoglucanase and xylanase activities are contributed by bacteria, and therefore, fiber associated bacteria are pivotal to the rumen digestion. These are divided into four groups on the basis of their association as free floating in the liquid phase, attached to the feed particles (firmly/ loose), rumen epithelium, protozoa, and fungi. Besides, these microbes have also diversified with respect to their functions. These carry out in degradation of plant ingredients. These have been classified as fiber (i.e., cellulose, hemicelluloses, and pectin)-degrading bacteria, lactic acid utilizers, acetogens, starch utilizers, etc. Starch and sugar digesters make up a significant part of the rumen bacterial population. Generally, high-producing dairy cows are fed diets containing more than 30 % starch and sugars; therefore, these bacteria are greatly needed. Even if a cow is on an all-straw diet, the fiber digesters still never account for more than 25% of the rumen bacterial population. Some microbes utilize the gaseous end products of the rumen to form methane and are grouped under archaea, while others are involved in converting the reducing power to useful end products as acetate, propionate, H2S, and butyrate. Always, there is an interest for non-methanogenic sinks to release H2 produced during fermentation. In the rumen, CH4 is produced by methanogens using H2 to reduce CO2 . A high-fiber- containing feed increases the population of Fibrobacter succinogenes , Ruminococcus albus , and Ruminococcus flavefaciens .The H2 and VFA production are also enhanced by high-fiber diet. Research on the effect of feeding ammonia-treated straw on rumen bacterial flora, showed that it stimulates the growth of Eubacterium ruminantium. Orpin et al. described seasonal variations of rumen bacteria in wild reindeer by demonstrating an increased cellulolytic Butyrivibrio fibrisolvens under severe dietary conditions in winter. Interestingly, non-cellulolytic bacteria were also detected as members of the fiber-associating consortia in the rumen Tannin-degrading bacteria have also been isolated from ruminants of wild origin identified as Selenomonas ruminantium as these consume tanniferous plants. Streptococcus bovis is present only when large amounts of starch or sugars are fed and pH is low. It produces lactic acid, a stronger acid than many of the other VFAs produced in the rumen. When conditions are favorable for Streptococcus bovis, it will grow explosively. This type of growth causes rumen acidosis. Megasphaera elsdenii requires lactic acid to grow that helps to clean up the rumen a bit and raise rumen pH, sustaining the growth of the acid-intolerant fiber digesters in the rumen.
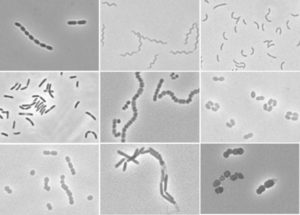
Rumen Bacteria
Anerobic Fungi
Fungi are the normal members of rumen ecosystem and are isolated under high fibrous feed. They are anaerobic and not found elsewhere 78% of rumen fungi associated with lignocellulosic component of plant material. These constitute the smallest in numbers and represent only about 20% of the rumen microbial biomass but are considered to be the most efficient fiber degrader. According to the recent classification system anaerobic fungi have been described under the order Neocallimastigales, class Neocallimastigomycetes in the phylum Neocallimastigomycota. The thallus may be monocentric (single reproductive body, i.e., sporangium from single zoospore) or polycentric (numerous sporangia from single zoospore). These forms are determined in the earliest stages of growth, soon after zoosporogenesis. The zoospores may be posteriorly uniflagellated or polyflagellated in both forms.The life cycle of anaerobic fungi is asexual, and no sexual stage has been described, as only mitotic nuclear divisions have been observed. It alters between a motile zoospore, encysted zoospore, and vegetative zoosporangial stage. Once released from zoosporangium, the motile zoospores move by chemotaxis to colonize the plant fibrous material and shed its flagella to get transformed into cyst(encystment). The cyst germinates by producing a germ tube, which ultimately gives rise to a rhizoidal system. In monocentric forms, the development may further be classified as endogenous or exogenous. These fungi differ from their aerobic counterparts in having hydrogenosomes as a site for energy production, instead of mitochondria.These fungi are not exclusively found in the rumen of the animals, but throughout the entire digestive tract. Moreover, these have also been isolated from the saliva and feces suggesting the presence of a resistant structure tolerant to the oxygen and desiccation. Formation of spores in the polycentric anaerobic gut fungi enhance their ability to survive and extend a possibility toward their use as direct-fed microbial. Anaerobic fungi have been isolated from a number of ruminant and non-ruminant herbivorous mammals.
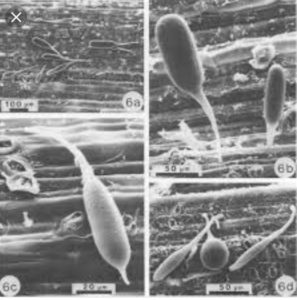
Rumen Anerobic Fungi
Protozoa
Rumen protozoa constitute about 50% of the viable biomass in the rumen, mass of rumen protozoa is equal to bacteria and 3% of rumen content. Majority are ciliates and few flagellates and are very motile. Unicellular organisms with 20–200 um size are not attached to feed particles. Protozoa are strictly anaerobic. They engulf bacteria and feed particles and digest carbohydrates, proteins, and fats. Two groups, i.e., holotrich and spirotrich protozoa, have been studied inside the rumen. Morphological studies have identified more than 250 species of ciliates living in the various ruminants. These ciliates play an important role in fiber digestion and the modulation of the fermentation profiles. The rumen protozoa produce fermentation end products similar to those made by the bacteria, particularly acetate, butyrate, and H2. Rumen methane bacteria actually attach and live on the surface of rumen protozoa for immediate access to H2. They utilize large amounts of starch at one time and can store it in their bodies. This may help to slow down the production of acids that lower rumen pH, benefiting the rumen. Rumen protozoa multiply very slowly in the rumen over 15–24 h as opposed to the bacteria. For this reason, the rumen protozoa hide out in the slower moving fiber mat of the rumen, so that these are not washed out before these have a chance to multiply. Low-roughage diets reduce the retention of fiber in the rumen and may decrease the number of protozoa in a cow’s rumen. The enzymatic profile of holotrich protozoa indicates that these have amylase, invertase, pectin esterase, and polygalacturonase in sufficiently large quantities for using starch, pectin, and soluble sugars as energy source The enzymes responsible for cellulose and hemicellulose degradation have also been reported in the holotrich protozoa, but the levels are very low compared to those present in the spirotrich protozoa. The rumen ciliates are proteolytic producing ammonia and amino acids as end products. Their N metabolism is based largely on the digestion of engulfed bacteria, although all rumen ciliates contain enzymes capable of digesting plant proteins. The utilization of these proportion of the total amount engulfed by the protozoa is excreted as amino acids. Degradation of protein or amino acids of microbial origin reduces the net yield of microbial amino acids available for intestinal digestion and hence, the net efficiency of microbial growth. Further, a large proportion of the protein incorporated the biomass of protozoa in the rumen may not be available for digestion in the intestines. Direct enumeration of protozoa in the rumen fluid entering the omasum concentration of a protozoa marker (i.e. phosphatidylcholine) in abomasal digesta, and measurement of the turn over rates of the protozoa all indicate a low outflow rate of ciliates from the rumen in relation to their concentration in the rumen. The synthesis of protozoa nitrogenous compounds and their breakdown in the rumen, therefore, contribute considerably to the N recycling within the rumen, as the apparent turnover of protozoa protein was calculated to be 0.67 of the total protein synthesized by protozoa.
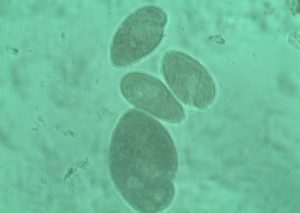
Holotrics
Methanogens
Methanogenic archaea, considered as the earliest organisms originated in our planet, are phyllogenetically widespread and represent three of the seven classes in the phylum Euryarchaeota. These rumen microorganisms utilize the H2 and CO2 produced by the protozoa, fungi, and bacteria from the catabolism of hexoses to produce CH4 and generate ATP which benefits the donors by providing an electron sink for reducing equivalents to minimize the partial pressure of H2 inside the rumen. These are comprised of about 70 species among 31 genera as reviewed in “Bergey’s Manual of Systematic Bacteriology” and “The Prokaryotes.” These microorganisms specifically possess three coenzymes, i.e., coenzyme F-420, involved in electron transfer; coenzyme M, involved in transfer of methyl groups; and a low molecular weight, oxygen-labile, heat stable coenzyme B, involved in the final reaction of CH4 production. Methanogens are found in a symbiotic association with rumen bacteria and protozoa. The establishment and maintenance of the stable population of methanogens is affected by the type of diet and level and frequency of feeding. The symbiotic relation between methanogens and ciliates may generate up to 37% of rumen CH4 emission. In the rumen, H2 is produced during plant cell wall degradation as an intermediate compound by cellulolytic bacteria (i.e.,Ruminococcus albus, Ruminococcus flavifaciens) and anaerobic fungi never gets accumulated, as it is rapidly utilized by methanogens in ruminants to produce CH4 and generate ATP.
Bacteriophages
Bacteriophages are one of the most important component of the rumen microbial community and are present typically at >109 particles per mL. These are specific for different bacteria present in the rumen. These are also considered to be obligate pathogens for the bacteria, as bacteriophages are capable of lysing bacteria. These phages help in bacterial mass turnover in the rumen, which may be considered not so useful for the animals on different feeding schedules but by lysing the bacterial cells, the bacterial protein is easily made available to the animals as a source of amino acids. The specificity of the bacteriophages for a particular rumen bacterium may be exploited for removal or killing by lysis of unwanted rumen bacteria from the ecosystem like Streptococcus bovis and methanogens. A little information is available on the genetic blueprint and gene functionality of archaea, particularly methanogenic, phages, but more are being discovered using electron microscopy and in vitro techniques. Researchers have reported siphophages that can infect methanogens (Methanobacterium, Methanobrevibacter, and Methanococcus spp.). Although these phages have not been isolated from the rumen. Despite the lack of knowledge, ruminant phage and their enzymes that are involved in lysing host cell represent significant opportunities for controlling both methanogen population and other community members in the rumen. In addition, it is possible to identify key host enzyme targets that are susceptible to inhibition by phage proteins understanding the life cycles of phage.
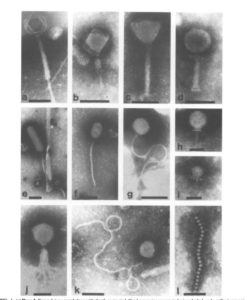
Bacteriophages
Author
Lonkare S.M.
Eastern Regional Station, National Dairy Research Institute, Kalyani, West Bengal
For any type of query related to animal husbandry, you can ask us by clicking ‘ask a question’ or ‘contact us’ option. For receiving a soft copy of any article, mail us at [email protected].
If you want to write an article or share your experience related to animal husbandry with other farmers, write your article and email it to [email protected].
If any student of veterinary science or veterinary doctor wishes to publish an article with us, then write your article and email at [email protected]. You can send your articles in Hindi, Punjabi, Urdu or English.

Thnku sir